The Applied Genomics Centre
For Infectious Diseases
For Infectious Diseases
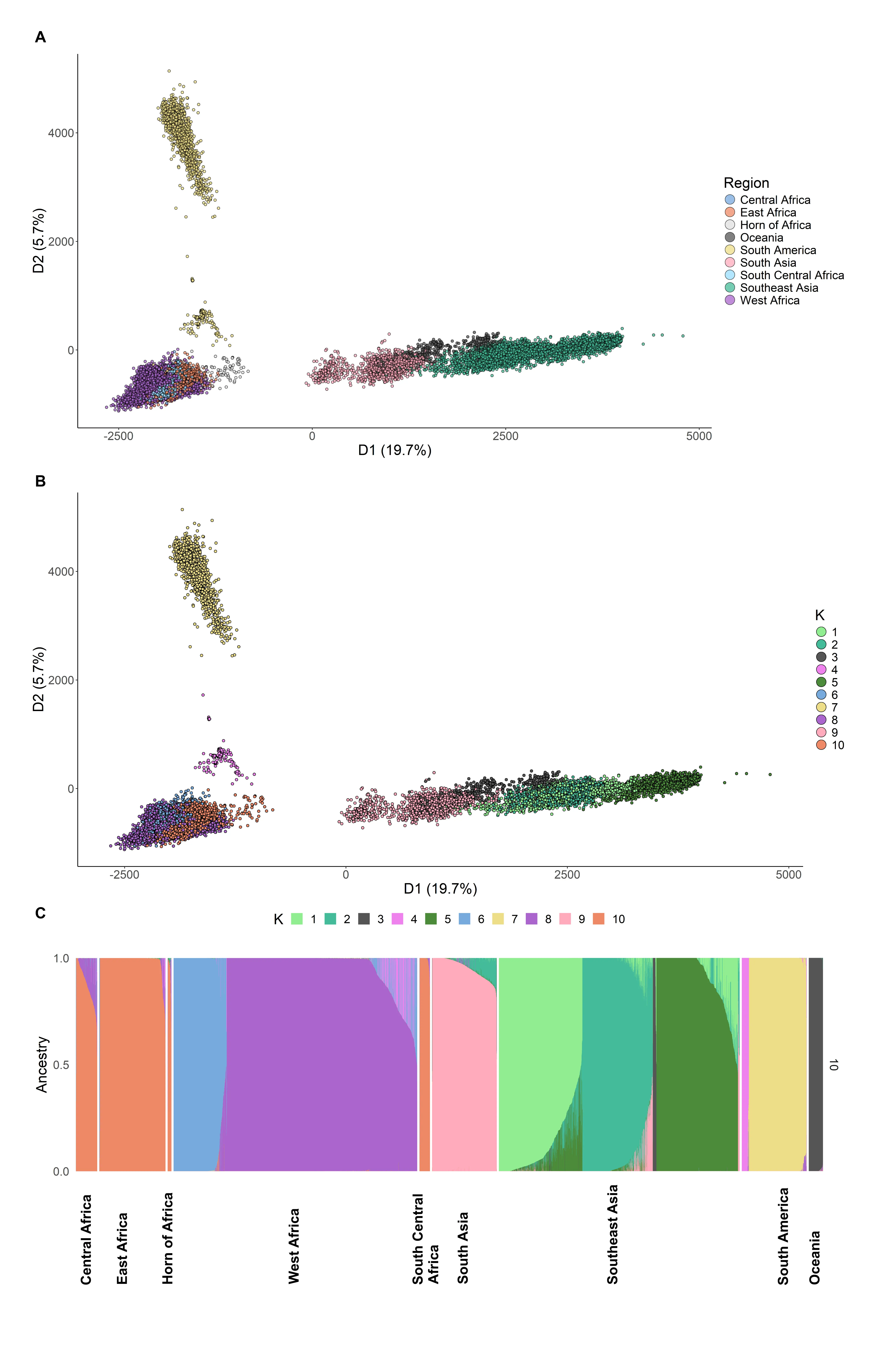
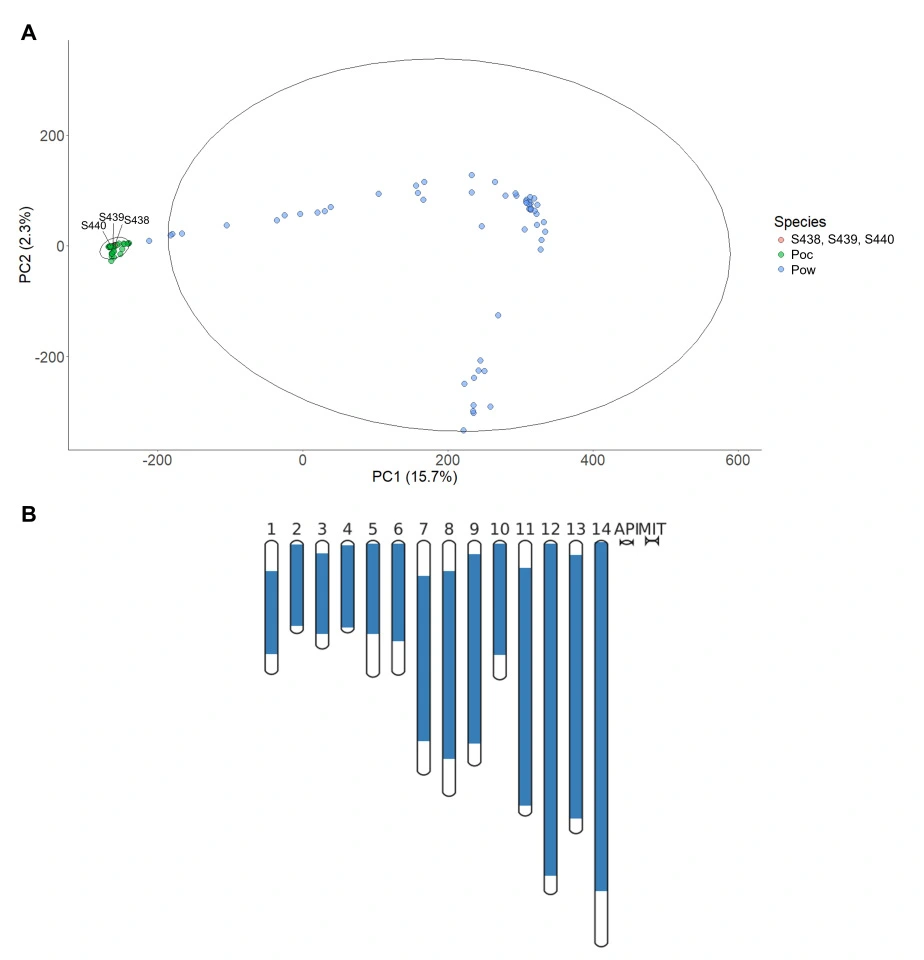

Large-Scale Genomic Analysis of Plasmodium falciparum Reveals New Insights into Geographic Diversity and Parasite Adaptation
A new study highlights the utility of sequencing and informatics in investigating cryptic malaria transmission
In April, we delivered our annual four-day LSHTM and online short course on infectious disease ‘omics.



Large-Scale Genomic Analysis of Plasmodium falciparum Reveals New Insights into Geographic Diversity and Parasite Adaptation
A new study highlights the utility of sequencing and informatics in investigating cryptic malaria transmission
In April, we delivered our annual four-day LSHTM and online short course on infectious disease ‘omics.